Uncovering the genetic architecture of inherited retinal disease in a consanguineous Iranian cohort

The Iranian IRD cohort under study consisted of 192 index patients and 622 family members (Fig. 1a, Supplementary Data 1). Consanguinity was reported for 76.1% of the cohort, with the majority corresponding to first cousin marriages (54.7%) (Fig. 1b). DNA from family members was available for 153/192 (79.7%) index cases. The gender balance was approximately equal, with 55.2% male and 44.8% female index patients (Fig. 1c). An overview of the phenotypes present in the cohort can be seen in Fig. 1d. The majority of index patients were diagnosed with RP (42.2%), followed by Leber congenital amaurosis (LCA, 17.7%) and CRD (12.5%). Syndromic IRDs were reported in 9.9% of the cohort. Phenotypic evaluation, if the participants cooperated, included color fundus photograph, infrared (IR) imaging, fundus autofluorescence (FAF), fluorescein angiography (FA), optical coherence tomography (OCT), visual field (VF), and electroretinography (ERG). This was complemented with demographic findings and pedigree information. The clinical results are accessible through the Iranian National Registry of Inherited Retinal Diseases (IRDReg®) software.
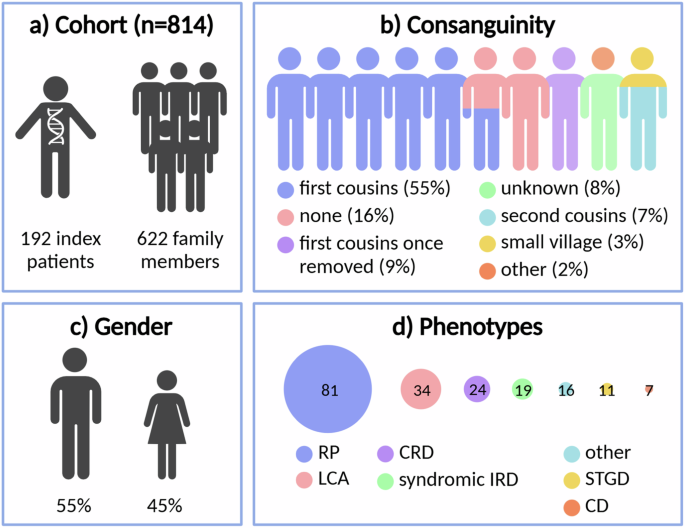
a Number of index patients and family members, b reported consanguinity, c gender, and d clinical phenotypes of the cohort. RP retinitis pigmentosa, LCA Leber congenital amaurosis, CRD cone-rod dystrophy, IRD inherited retinal disease, STGD Stargardt disease, CD cone dystrophy, other: all phenotypes not included in the remaining categories. Created in https://BioRender.com.
Autozygosity-driven WES-based testing elucidates the molecular cause of IRD in 72.9%
WES and subsequent analysis of single-nucleotide variants (SNVs) and copy number variants (CNVs) in a panel of known IRD-associated genes (RetNet panel) uncovered the (likely) cause of IRD in 140/192 (72.9%) index patients (Fig. 2a). These included 132 (68.7%) solved (one heterozygous class 4/5 variant for AD disease, one hemizygous class 4/5 variants for XL disease, one homozygous or two heterozygous class 4/5 variants in trans for AR disease) and 8 (4.2%) likely solved cases (two heterozygous class 4/5 variants without segregation data or a heterozygous class 3 variant in trans with a heterozygous class 4/5 variant for AR disease). Another 29 patients (15.1%) were considered to have an uncertain diagnosis, based on the presence of variants of uncertain significance (VUS) (class 3 variants for AD or XL disease, two heterozygous class 3 variants in trans, a heterozygous class 4/5 variant together with a heterozygous class 3 variant without segregation data, two heterozygous class 3 variants without segregation data or a homozygous class 3 variant for AR disease). No plausible cause of IRD was found for the remaining 23 patients (12.0%). The highest (likely) solved diagnostic yields were found in patients with ‘other’ (87.5%), Stargardt disease (81.8%) and LCA phenotypes (79.4%) (Supplementary Fig. 1). These are followed by CRD (79.2%), syndromic IRD (73.7%), RP (65.4%) and CD (57.1%). ‘Other’ phenotypes are those different from the mentioned categories and include patients with the following diagnoses: achromatopsia (n = 4), choroideremia (n = 4), bestrophinopathy (n = 3), Usher syndrome (n = 2), clumped pigmentary retinal degeneration (n = 1), early RCD with macular dystrophy (n = 1), Goldmann-Favre syndrome (n = 1), optic atrophy (n = 1) and retinitis pigmentosa or cone-rod dystrophy (n = 1).
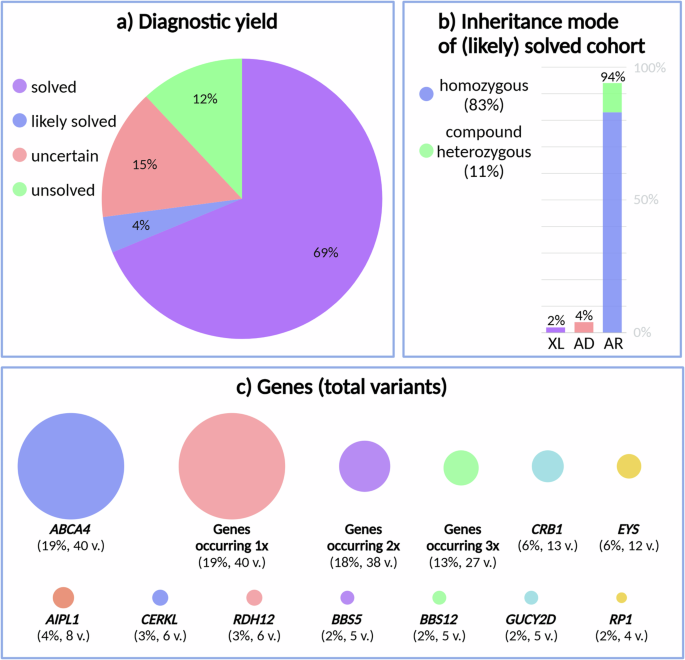
a Diagnostic yield, b identified inheritance mode in the (likely) solved cohort, and c genes in which variants were found (total number of variants: every occurrence counted once). XL X-linked, AD autosomal dominant, AR autosomal recessive, and v. variants. Created in https://BioRender.com.
Within the patient cohort that was considered to be (likely) solved or has an uncertain molecular diagnosis, the majority showed homozygosity for the identified variants (68.4%), followed by compound heterozygosity (23.4%), heterozygosity (6.2%) and hemizygosity (1.9%). For the (likely) solved patients only, AR inheritance was identified in 94.0%, with 82.8% corresponding to homozygous variants and 11.2% to compound heterozygous variants (Fig. 2b). Only 4.5% of (likely) solved patients were found to harbor variants in IRD genes associated with AD inheritance. Finally, variants in genes associated with XL inheritance were identified in 1.5% of the (likely) solved patients. Approximately 82.8% of variants found in patients with a known consanguineous background were located in an ROH.
The vast majority of identified variants in the IRD cohort are unique and novel
A total of 209 variants were identified across the solved, likely solved and uncertain cohorts, encompassing 78 distinct genes associated with IRD (Fig. 2c). Strikingly, the majority of these variants were unique (Fig. 3a). Among the cohort studied, the most frequently observed variant was c.5882 G > A, p.(Gly1961Glu) in the ABCA4 gene, occurring 7 times. Additionally, a CERKL variant (c.769 C > T, p.(Arg257Ter)) was found three times, followed by 14 other recurring variants, each appearing twice. For the (likely) solved IRD patients, 54.9% of variants were classified as pathogenic, 41.9% as likely pathogenic and 3.2% as VUS.
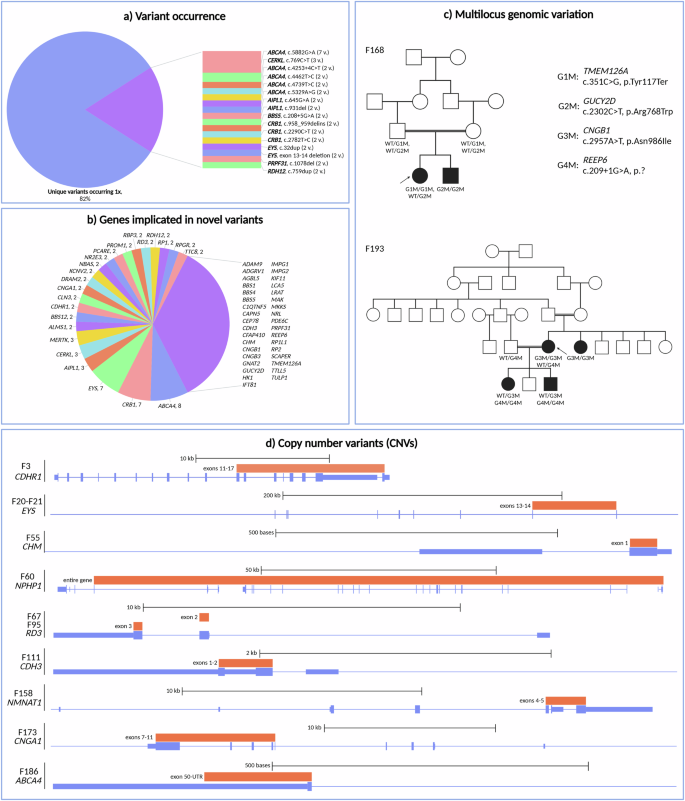
a Variant occurrence, b genes in which novel variants were found and the number of novel variants per gene (genes in which only one novel variant was found are grouped together at the right), c pedigrees of two families in which multilocus genomic variation was identified and d overview of the locations of the CNVs that were found in the cohort. WT wild type. Created in https://BioRender.com.
Out of the 187 unique variants, approximately half (52.9%) were novel. These novel variants were found in 58 distinct genes, with the largest number found in ABCA4 (n = 8; representing 26.7% of all ABCA4 variants), CRB1 (n = 7; 70.0% of CRB1 variants), EYS (n = 7; 70.0% of EYS variants), AIPL1 (n = 3; 50.0% of AIPL1 variants), CERKL (n = 3; 60.0% of CERKL variants) and MERTK (n = 3; 100.0% of MERTK variants). The other implicated genes are shown in Fig. 3b. The vast majority of novel SNVs (92.5%) were not present in the Iranome database, containing 800 Iranian healthy individuals from eight ethnic groups21. Only 7.5% could be found in Iranome, with a maximum frequency of five heterozygotes. Genes in which only novel variants were discovered and the number of variants are listed in Supplementary Fig. 2. Out of the 14 variants recurring twice (see above), six variants were novel and occurred in two patients each (AIPL1 c.931del, p.(Arg311Alafs*17); BBS5 c.208+5 G > A, p.?; CRB1 c.958_959delinsACATGTGAG, p.(Val320Thrfs*4); CRB1 c.2782 T > C, p.(Cys928Arg); PRPF31 c.1078del, p.(Arg360AlafsTer3) and RDH12 c.759dup, p.(Phe254LeufsTer19)).
Distribution of SNVs and CNVs in IRD genes shows 95% of SNVs and 5% of CNVs
The vast majority of the 187 identified variants (94.7%) in known IRD genes were SNVs: 43.9% missense variants, followed by 18.7% frameshift, 16.6% nonsense, 12.3% splice, 1.6% in-frame and 1.6% synonymous variants. A distribution of the most frequent genes in which SNVs were found, can be seen in Fig. 2c. Following SNV analysis, WES data from all patients was examined with ExomeDepth to identify potential CNVs. CNVs within known IRD genes were detected in 11/192 patients (5.3%) (Table 1). Notably, all CNVs were deletions (100.0%), identified in nine different IRD genes. Among these, an identical deletion in EYS was observed in two families, while two distinct deletions in RD3 were found in two different families. Most of the deletions were out-of-frame (80.0%). Single-exon deletions were identified in 40.0%, multiple-exon deletions in 50.0%, and a complete gene deletion in 10.0%. Additionally, 7/10 unique deletions (70.0%) were novel and located within the ABCA4, CDH3, CDHR1, CHM, CNGA1 and RD3 genes. The CNV positions relative to the genes are shown in Fig. 3d.
WES dissects distinct phenotypes resulting from multilocus genomic variation
In two families, more than one genotype segregated with disease, illustrating multilocus genomic variation. The pedigrees are shown in Fig. 3c. In F168, a homozygous TMEM126A variant (c.351 C > G, p.(Tyr117Ter)) (class 4) was identified in the index case. Segregation analysis revealed heterozygosity of this variant in both parents, but absence in an affected sibling. Further analysis of the affected sibling uncovered a homozygous GUCY2D variant (c.2302 C > T, p.(Arg768Trp)) (class 5), found in a heterozygous state in the index case and in both parents. The index case displayed optic atrophy, while the affected sibling was diagnosed with LCA, a distinct phenotype.
Another example of multilocus genomic variation was observed in F193, a family with RP. A homozygous CNGB1 variant (c.2957 A > T, p.(Asn986Ile)) (class 4) was found in the index patient and her affected sibling. The affected children of the index case were only heterozygous carriers, however. Upon further investigation, a homozygous REEP6 variant (c.209+1 G > A, p.?) (class 4) was identified in both affected children, with both parents being heterozygous carrier of this REEP6 variant.
Rare genotype-phenotype correlations
Two heterozygous NBAS variants (c.1194_1195del, p.(Asp398GlufsTer29) and c.5671 G > A, p.(Asp1891Asn)) were identified in the index patient of F34, diagnosed with LCA with no syndromic features. Available clinical data is presented in Supplementary Fig. 4. NBAS variants are associated with infantile liver failure syndrome (OMIM #616483) and short stature, optic nerve atrophy and Pelger-Huët anomaly (OMIM #614800). Additionally, one patient with NBAS variants has been described, presenting with CD, optic atrophy and Pelger-Huët anomaly but no other symptoms22. To our knowledge, no other non-syndromic cases have been previously reported in literature.
Two RP patients in the cohort were found to carry possible disease-causing DRAM2 variants (c.132-6 T > C, p.? and c.314 G > T, p.(Gly105Val)). The clinical diagnosis of one patient (F8) was refined to RP sine pigmento with macular involvement. The macular aspect of this phenotype could be in line with the DRAM2-associated AR retinal dystrophy with early macular involvement23. However, the phenotype of F97, end-stage RP without mention of macular problems, is atypical for DRAM2-associated IRD (see Supplementary Figs. 3 and 6 for available clinical data).
A homozygous CDH3 deletion of exons 1–2 was identified in F111, a patient with early RCD with macular dystrophy. CDH3 variants, however, are associated with congenital hypotrichosis with juvenile macular dystrophy (HJMD) (OMIM #601553)24. Symptoms of this disease are childhood-onset and progressive macular dystrophy combined with sparse scalp hair (hypotrichosis). Interestingly, CDH3 variants have also been shown to cause CRD, with the hypotrichosis confined to hypoplastic nails only25. In a family of Druze origin, three siblings with a CDH3 variant were additionally identified, their phenotype was described as RP, without skin or hair abnormalities26. Our findings could confirm this rare association of biallelic CDH3 variants with non-syndromic RP. Clinical reexamination to exclude extra-ocular symptoms was, however, not possible. Available clinical data is included in Supplementary Fig. 7.
Biallelic variants in CEP78 are typically associated with AR CRD and hearing loss (CRDHL) (OMIM #617236). Two causal variants (c.356 C > T, p.(Ser119Leu) and c.515 T > G, p.(Ile172Arg)) in this gene were found in homozygous state in two families in this Iranian cohort. In F141, with a clinical diagnosis of CRDHL (see Supplementary Fig. 8), c.515 T > G, p.(Ile172Arg) was identified, which has already been found in patients with CRDHL27. In F40, however, where the novel CEP78 variant c.356 C > T, p.(Ser119Leu) was identified, the phenotype was determined as RP without reported hearing impairment (see Supplementary Fig. 5). Other rare CEP78 cases have been described in literature: two siblings with non-syndromic RP28 and a patient with CRD without hearing loss29.
Phenotypic heterogeneity in genes associated with syndromic IRD
CLN3 variants can cause neuronal ceroid lipofuscinosis, a syndromic IRD (OMIM #204200), as well as isolated retinal dystrophy30,31,32,33. Both phenotypic presentations were also found in the studied cohort: syndromic IRD in F45 (c.1274_1275del, p.(Leu401ProfsTer7)) and isolated RP in F48 and F70 (c.1213 C > T, p.(Arg405Trp) and c.1056 G > A, p.(Gln352 = )). The variant in F48 has been described as a cause of non-syndromic retinal dystrophy in literature30,31,34,35. However, two siblings homozygous for this c.1213 C > T, p.(Arg405Trp) variant, initially diagnosed with isolated retinal dystrophy, developed neuronal symptoms in their third decade36. This variant is also homozygously present in F48, therefore monitoring of any late-onset neuronal symptoms will be advised. The other two identified CLN3 variants (c.1056 G > A, p.(Gln352 = ) and c.1274_1275del, p.(Leu401ProfsTer7)) have not yet been reported.
link